Multivariate analysis of of covariance (MANCOVA) is used to test the statistical significance of the effects of one or more independent variables on a two or more dependent variables, after controlling for covariates.
MANCOVA, like other statistical tests, requires data to meet several important assumptions in order to get meaningful results. The assumptions are the same for MANOVA, with additional requirements relating to covariates. This leads to a laundry list of assumptions, which can make the analysis a bit overwhelming for beginners. However, if you’re using software (like SPSS), testing for assumptions becomes less of a challenge, and is the main reason why MANCOVA is very rarely performed by hand.
What are the MANCOVA Assumptions?
The basic assumptions are:
- Level of Measurement: Independent variables should be categorical; Dependent variables should be measured at the interval level or above (i.e. scale variables or continuous variables). Covariates have more flexibility and can be continuous, dichotomous, or ordinal.
- Independence of observations: Observations must be independent of all other observations; If your data has been drawn using a random sampling method, this assumption is met.
- The dependent variables should be normally distributed within groups.
- Homogeneity of variances must be met for each dependent variable. In other words, the variances in the difference groups must be the same.
- Homogeneity of covariances: The intercorrelation matrix between dependent variables must be equal for all levels of the independent variable.
- There must be a significant linear relationship between the dependent variable and the covariate [1].
- The slope of the regression line must be the same in each group.
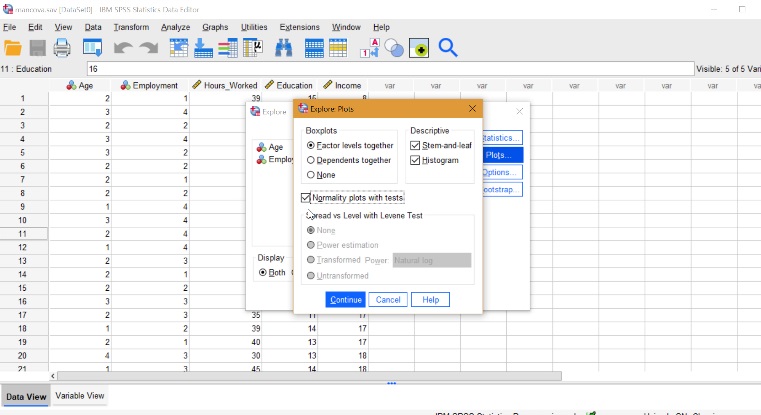
If you want to test for these assumptions, use software: See: How to Run a MANCOVA in SPSS .
Effect of violating MANCOVA assumptions
If your data severely violates assumptions, your p-values become meaningless and you open your research up to higher risk of Type I and Type II errors.
Violating some assumptions, like multivariate normality, leads to minimal effect on your analysis; The F test is very robust to deviations from normality. However, the assumption of normality should still be taken into consideration to make sure your data doesn’t deviate entirely from the assumption. In general, if your kurtosis is greater than 0, the F statistic will be too small to reject the null hypothesis [2].
Other violations are much more problematic, including violating the assumption of independence, which leads to an increased likelihood of type I error in the F statistic. It also interferes with standard errors of means and inferences about them.
Violating the assumption of homogeneity of variance-covariance matrices leads to a whole host of problems, including increasing the likelihood of Type I and Type II errors. If you have an unbalanced design, with a ratio of 1:2 or more, Type I error rates increase dramatically, even with mild heterogeneity [3].
References
[1] Siri, F. et. al. (2018). The Arts and The Brain. Retrieved October 12, 2021 from: https://www.sciencedirect.com/topics/medicine-and-dentistry/multivariate-analysis-of-covariance
[2] ANOVA/MANOVA.
Retrieved October 12, 2021 from: https://sites.oxy.edu/lengyel/m150/textbook/stanman.html#assumptions
[3] MANOVA and MANCOVA. Retrieved October 12, 2021 from: