What is a Greedy Algorithm?
The greedy algorithm is one of the simplest algorithms to implement: take the closest/nearest/most optimal option, and repeat. It always chooses which element of a set seems to be the best at the moment. It never changes its mind at a later point.
Greedy algorithms are “top-down”, which mean that the algorithm makes one greedy choice and then another, reducing large problems to smaller ones. The idea is that by choosing the tastiest (most optimal) element at any moment, the overall system will eventually be optimized. Most problems cannot be optimized by a greedy algorithm, but it does work for some cases (like greedy matching).
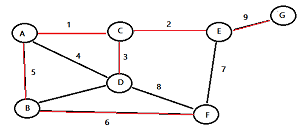
Greedy Matching Algorithm
The goal of a greedy matching algorithm is to produce matched samples with balanced covariates (characteristics) across the treatment group and control group. It can generate one-to-one or one-to-many matched pairs sampled without replacement. Sampling without replacement means that once a person has been matched, they cannot be used in another match.
Greedy nearest neighbor is a version of the algorithm that works by choosing a treatment group member and then choosing a control group member that is the closest match. For example:
- Choose the participant with the highest propensity score (a propensity score is the probability of being assigned to the treatment group).
- Select a control group member with the closest propensity score to the person picked in Step 1.
- Choose a second treatment group member (in this example, with the next highest propensity score rank), match the second participant.
- Repeat the process until all participants are matched.
You don’t have to start with the participant with the highest rank propensity score. Other options are:
- Lowest to highest score.
- Best possible match: a pair is chosen with the closest scores. They are removed from the set, then another pair is chosen with the next closest scores, and so on.
- Random selection: pairs are chosen at random using a random seed.
- Caliper matching: a maximum caliper distance is set for the matches. A caliper distance is the absolute difference in propensity scores for the matches. As a maximum value is being set, this may result in some participants not being matched.
Disadvantages
Greedy nearest neighbor matching may result in poor quality matches overall. The first few matches might be good matches, and the rest poor matches. This is because one match at a time is optimized, instead of the whole system. An alternative is optimal matching, which takes into account the entire system before making any matches (Rosenbaum, 2002). When there is a lot of competition for controls, greedy matching performs poorly and optimal matching performs well. Which method you use may depend on your goal; greedy matching will create well-matched groups, while optimal matching created well-matched pairs (Stuart, 2010).
Greedy Matching Programs
- In R: Run MatchIt. The default nearest neighbor matching is greedy matching.
- In SAS: Several macros are available for one-to-one matching including Parson’s macro and Kosanke & Bergstral’s gmatch.
References:
Austin, P. (2014). A comparison of 12 algorithms for matching on the propensity score. Retrieved February 3, 2017 from: Stat Med. 2014 Mar 15; 33(6): 1057–1069.
ROSENBAUM, P. R. (2002). Observational Studies, 2nd ed. Springer, New York. MR1899138
Stuart, E. (2010). Matching methods for causal inference: A review and a look forward. Stat Sci. 2010 Feb 1; 25(1): 1–21.